Scientist pulls protein through microscopic hole
A Danish researcher has developed a new method for studying proteins. By pulling the protein through a tiny hole known as a nanopore, he can expose important properties of the protein.
Humans, giraffes, woodlice or snakes. All living organisms contain proteins, and they are essential building blocks of life.
But what exactly do the tiny proteins look like?
A Danish scientist has discovered a new way to reveal certain details about protein molecules.
”One of the advantages of using this technique is that it is extremely sensitive, which enables us to detect very small characteristics of the proteins,” says Christian Bech Rosen, a PhD fellow at the iNANO Centre at Aarhus University.
Phosphorylation occurs everywhere in Nature – it is one of Nature’s most common ways of controlling protein function. In some proteins, the phosphorylation functions as a kind of on/off switch. That’s why it is important to learn about how proteins are phosphorylated.
The details of Rosen’s new technique are published in the journal Nature Biotechnology.
Protein pulled through channel
The novelty of the new study is not that it makes it possible to determine important information about protein composition. That’s something scientists have been able to do for some years now.
The novelty lies in the method used to find this information.
In short, Rosen has managed to pull single proteins, one at a time, through a microscopic channel in a membrane. By observing what happens the exact moment the protein goes through the channel, he can analyse important structural details of the protein, including where a so-called phosphate group is located.
Phosphate groups are important for life
Phosphate groups are small chemical moieties that play an important role in life. The body’s metabolism, for instance, is dependent on these phosphate groups.
The process in which a phosphate group binds to a protein is known as phosphorylation. Rosen’s method offers a new way of examining where a protein is phosphorylated.
”Phosphorylation occurs everywhere in Nature – it is one of Nature’s most common ways of controlling protein function. In some proteins, the phosphorylation functions as a kind of on/off switch,” says Rosen.
”That’s why it is important to learn about how proteins are phosphorylated.”
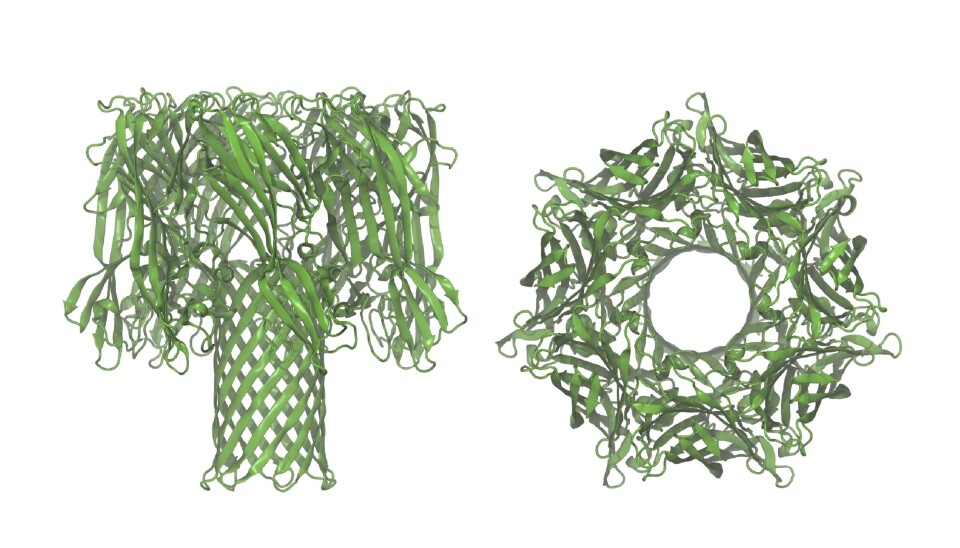
In his new study, Rosen used a well-known instrument consisting of a vessel with brine, which is separated at the centre by a lipid membrane. A so-called nanopore was then inserted into the membrane, which acts as a nanoscale channel that is the only place a protein can move from one side of the membrane to the other.
Rosen then applied a potential difference between the two sides of the vessel, allowing a current – a flow of ions from the salt in the water – to pass through the channel.
Protein modified with DNA
However, the protein cannot translocate the nanopore by itself. In order to get the protein to move through the pore to the other side of the vessel, the researcher first made minor alterations to the protein.
“The protein will not by itself pass through the pore. It requires something to pull it through.”
Rosen therefore modified the protein with a tiny DNA strand, which is electrically charged and could therefore pull the protein through the nanopore to the other side of the membrane in the vessel.
At the exact moment the protein slipped through the pore in the membrane, it blocked the ionic current from going through the pore.
“When the protein is pulled through the nanopore, the pore becomes partially blocked, and that’s something we can read off as a brief change in the ionic current,” says Rosen.
Ionic current indicates phosphate groups
The researchers used the information about the changing current in the nanopore to determine exactly where the phosphate groups are attached to the protein.
“Let’s say that a protein can be phosphorylated in two locations. If it is phosphorylated in one location, you get a current signature that looks a certain way, and if the protein is phosphorylated in another location, you get a different signature,” says Rosen, adding that the technique is not yet fully developed:
”We do not yet have a finished product for studying protein phosphorylation, but hopefully this will be the starting point for future results where nanopore technology is used in this way,” he says.
“It is remarkable that we can identify a tiny phosphate group in a huge and complex protein based on how it blocks the nanopore. This opens up for the possibility to study many other small – but essential – modifications in protein structures using nanopore technology.”
Hagan Bayley, a professor at University of Oxford and co-author of the new study, says:
“Our paper is the very first attempt to examine posttranslational modification with nanopores, and further substantial developments can be expected.”
--------------
Read the Danish version of this article at videnskab.dk