Researchers' Zone:

The medicines of the future are hiding in Nature's treasure trove
Researchers at the University of Copenhagen are studying bioactive molecules in fungi to find the medicines of tomorrow.
Analysis shows that, by 2050, 10 million people will die annually of infections from multidrug-resistant bacteria, and that more than 630 million people will be diagnosed with the lifestyle disease type 2 diabetes.
In order to break this trend and address the huge health challenges facing society, new medicines need to be developed.
In the Natural Product Research group at the Department of Drug Design and Pharmacology at the University of Copenhagen, we delve into Nature's treasure trove to find the compounds that have the potential to become the medicines of the future.
In the following, I will look at how we natural product researchers navigate Nature's gigantic collection of molecules to find those specific ones that have the potential to become the drugs of the future.
'Holy bark' cured malaria
Traditionally, Nature has been our main source for both the treatment of diseases – in the form of traditional medicines and stimulants such as coca leaves – and a wide range of poisons for hunting purposes.
In addition to being enriched with an abundance of bioactive substances, nature contains a fabulous chemical diversity that yields medicines with new treatment mechanisms and thus a possible solution to the increasing development of resistance we see among bacteria, for example.
One of the best examples of nature's creativity is to be found in the treatment of malaria, a fever-like disorder caused by an infection from parasites of the Plasmodium genus, which is transmitted via mosquito bites.
In the Amazon jungle, the bark of the cinchona tree (cinchona bark) has traditionally been used to treat fever, and it was described in the 1600s as 'holy bark' by missionaries in South America.
Based on this ethnopharmacological knowledge, the drug quinine was isolated from cinchona bark and used to treat malaria in 1820 – and is still in use to this day.
When nature is the source of future medicines
During the Vietnam War, the North Vietnamese suffered heavy losses due to malaria infections, many of which were resistant to quinine and its analogues, leading to their ally, China, to launch a large-scale campaign in 1967 to find new drugs to treat malaria.
Again, they turned to traditional medicine and investigated Chinese medicinal plants used to treat fever.
This led to the isolation of artemisinin from Artemisia annua (sweet wormwood), which to this day is the preferred drug for the treatment of malaria due to having fewer side effects and less resistance development than quinine analogues.
This example clearly shows why nature is such an important source of the medicines of the future.
The solution to the challenges of side effects and resistance development was not to make synthetic analogues of quinine but rather to discover another and markedly different molecule from Nature's treasure trove.
Despite the fact that filamentous fungi (moulds), unlike plants, are not widely used in traditional medicine, they have nevertheless been a source of one of the world's most important medicines, penicillin.
Since the discovery of penicillin, a large number of drugs have been isolated from filamentous fungi, including the cholesterol-lowering agent lovastatin, antibiotics such as cephalosporin and immunosuppressants such as cyclosporin A, which are used in organ transplantations, among other things.
When filamentous fungi play hide and seek
If we as natural product researchers are to find new bioactive molecules, we must first collect the biomass that we want to investigate.
This can be anything from plants to filamentous fungi and marine organisms, and the selection criteria will normally depend on the diseases to be treated.
One of the promising sources we work with in our research group is endophytic fungi, which can be defined as fungi that live inside plants without harming them.
Fungi can sometimes have a beneficial effect on plants in the form of providing defence compounds against insects, for example, in exchange for nutrition from the plant.
Before we can start examining endophytic fungi, they need to be extracted from the plant and into the laboratory. We do this by luring them out with lots of nutrition.
The procedure consists of sterilising the surface of the plant in order to kill all the microorganisms found on the outside and then making some incisions in the plant material from which the endophytes can emerge.
When the endophytes emerge out of the plant, we can cultivate them in the lab and examine the many compounds they produce.
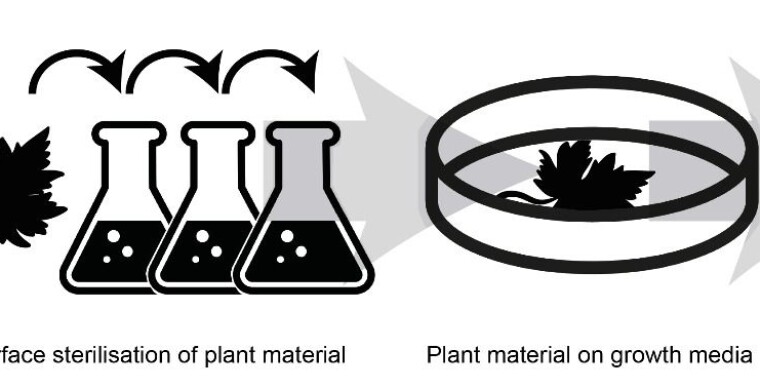
Fungal schnapps in the lab
A single filamentous fungus can produce a complex mixture of secondary metabolites – molecules that are not essential for the fungus’ survival but which give them an advantage in nature – with up to 100 different substances.
Often, however, the genes that encode the multiple substances are not active under the standard conditions in laboratories, so the fungus is grown on a variety of media in order to activate as many genes as possible.
Once the fungus has grown for 7 to 14 days, it is extracted with organic solvents. It sounds technical – and it is – but it is comparable to making a fungal schnapps (which, however, is not recommended for your birthday party).
To find out if the fungus contains secondary metabolites with interesting activity, we test the extract against a range of relevant bacteria, cancer cells or specific enzymes involved in the disease we want to treat.
From complex mixtures to a single substance with biological activity
One of our most important tasks is to find the exact compound or compounds in the fungus that give rise to the observed activity we are seeking for the future medicines.
In our research group, we use a method that allows us to point out the active compound(s) directly in the aforementioned complex fungal snaps – this is what we call a bio-chromatogram.
The method is to separate all the fungus’ constituents based on their polarity (whether they are soluble in water or oil) using so-called high-performance liquid chromatography (HPLC).
This is done by using solvents that push the metabolites through a column filled with a porous material. This allows the metabolites to be divided in 20 to 50 minutes, depending on the complexity.
At the end of the column in which the metabolites are separated, a detector provides us with a signal so we can see when the substances are being pushed through the column.
As the metabolites are pushed through the column, they are collected in 80 to 160 equal fractions, and by testing these for the desired activity, we can see which metabolites are active in the complex mixture.
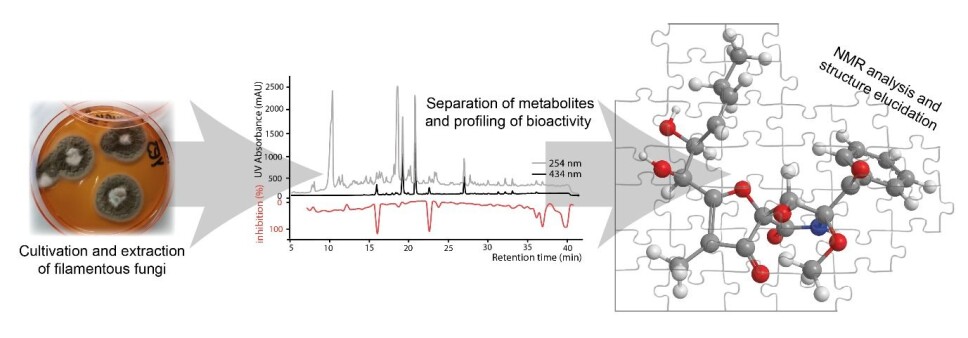
Using various methods, including HPLC, we isolate the interesting compounds to determine the biological activity and identify the chemical structure.
Solving the structure - a 3D puzzle without a predefined model
Once we have isolated a molecule with biological activity, the next challenge is to determine the structure – how the atoms are connected to each other.
For this, nuclear magnetic resonance spectroscopy is the preferred method. Here we take advantage of the fact that the nuclei of certain atoms – including hydrogen, carbon and nitrogen – behave like small magnets when placed in a very strong magnetic field.
Depending on the magnetic environment in which the individual nuclei are located, they will behave differently in the strong magnetic field. We use these differences to generate a series of 1D and 2D spectra, where we can both see the differences and how the individual atoms interact with each other.
Solving a structure can be compared to solving a 3D puzzle where, based on the generated NMR spectra, we define the size, shape and number of the pieces and then put them together.
Do future medicines lie at the extremes of nature?
There is no doubt that society is facing major health challenges that need new medicines.
However, as shown in the example of antimalarial drugs, new medicines with alternative active mechanisms are most often found in completely different places than known compounds.
A key area of our pharmaceutical research is, therefore, to examine filamentous fungi from extreme environments, such as those with high temperatures, extreme salt concentrations or high acid content.
The idea behind this is that filamentous fungi from these environments must be quite unique and thus potentially contain unique molecules that can be developed for the drugs of the future.
With the continued growth in resistance and increased incidence of cancer, there is a growing need to delve into nature's treasure trove for tomorrow's bioactive molecules.
So, remember to appreciate the little green spots on your old bread before you throw it out – you never know what wonders they may contain!
Translated by Stuart Pethick, e-sp.dk translation services. Read the Danish version at Videnskab.dk's Forskerzonen.




